October 3, 2013
Seat reservations in the developmental stage
Gašper Tkačik publishes paper on positional information of genes in embryos with Nobel laureate Eric Wieschaus
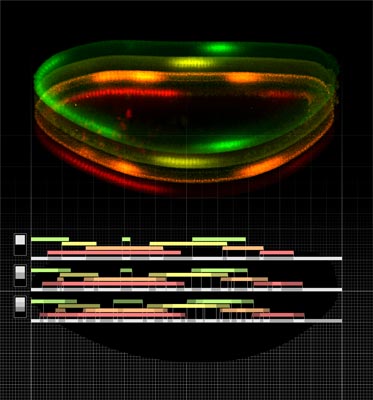
For a head to become a head, and a tail a tail, cells in a developing embryo need to make decisions and adopt proper fates depending on where in the embryo they are located. How precisely cells can “find out” which position they have in the embryo is the subject of a study appearing this week in the Proceedings of the National Academy of Sciences (PNAS) authored jointly by Julien Dubuis and Gašper Tkačik, and collaborators Thomas Gregor, Nobel laureate Eric Wieschaus, and William Bialek. The study is published as an Inaugural Article on the occasion of the election of William Bialek to NAS last spring, and is a result of a continued collaboration with Gašper Tkačik, now Assistant Professor at IST Austria, who did his PhD in Physics under the supervision of Bialek at Princeton University.
Cells in a developing embryo have no direct way of measuring their physical position; however, this “positional information” is essential for correct development. While it has been known that positional information is provided by gene expression levels of special developmental genes expressed along the long axis of the embryo (known in the fruitfly Drosophila melanogaster as gap genes), this knowledge has remained qualitative. It was unknown “how much” positional information gene expression levels encode, or even how such a question could be asked with mathematical rigor and estimated from experimental data. In their work, the researchers took up the task of putting “positional information” onto a firm basis using information theory. They asked what the level of expression of a particular gap gene in a cell tells this cell about its position, how much information gap genes carry, either singly or in their joint expression pattern, and how much of this information the embryo actually uses.
The results published in PNAS show that the patterns of gap gene expression provide enough information to specify every cell’s position with a relative error as low as 1%. This corresponds to the observed precision of subsequently formed pattern and structure elements in the embryo, like the cephalic furrow, for which gap genes serve as inputs. This quantitatively demonstrates that all the required patterning information along the long embryo axis can be encoded by the gap genes. The expression pattern of each gap gene carries 2 bits of information, which is more information than in the previously held textbook view, where gap genes define broad ON or OFF domains of expression separated by sharp boundaries; the traditional view would provide at most 1 bit of information per gene. Comparing how much positional information the gap genes carry to the theoretical maximum, the researchers find that the gap genes in the fruitfly form an astonishingly efficient code for position. This result suggests an interesting link to evolution: did natural selection favor genetic circuits that are optimal in an information-theoretic sense?



